Model#
- class cosmolayer.cosmosac.Model(min_sigma, max_sigma, num_points, area_per_segment, averaging_radius, f_decay, sigma_0, merge_profiles, temperature_exponents, interaction_matrix_generator)[source]#
Immutable configuration for a COSMO-SAC model variant.
Bundles all model-specific parameters into a single object, ensuring that components and mixtures are always created with consistent settings. Use the pre-built
CosmoSac2002ModelandCosmoSac2010Modelsingletons for standard calculations, or construct a custom instance for research purposes.- Parameters:
min_sigma (float) – Minimum screening charge density in e/Ų.
max_sigma (float) – Maximum screening charge density in e/Ų.
num_points (int) – Number of discrete points in the sigma grid.
area_per_segment (float) – Reference surface area of a single segment in Ų.
averaging_radius (float) – Effective radius for distance-weighted sigma averaging in Å.
f_decay (float) – Decay factor for exponential distance weighting in the sigma averaging procedure.
sigma_0 (float or None) – Standard deviation of the Gaussian probability of a segment to form a hydrogen bond in e/Ų. Set to
Noneto disable hydrogen-bond splitting (all surface area is assigned to the NHB class).merge_profiles (bool) – Whether segment-group profiles (NHB, OH, OT) should be merged into a single distribution by default when computing probabilities.
temperature_exponents (tuple[int, ...]) – Exponents applied to the temperature in the interaction energy expression. Must have the same length as the tuple returned by
interaction_matrix_generator.interaction_matrix_generator (Callable) – Function
f(temperature) -> tuple[NDArray, ...]that produces the dimensionless interaction energy matrices ΔW/(RT) at a given temperature.
Examples
Using a pre-built model to create a component:
>>> from importlib.resources import files >>> from cosmolayer.cosmosac.model import CosmoSac2002Model >>> path = files("cosmolayer.data") / "C=C(N)O.cosmo" >>> component = CosmoSac2002Model.create_component(path.read_text()) >>> component.area 97.34554...
Inspecting model parameters:
>>> CosmoSac2002Model.area_per_segment 7.5 >>> CosmoSac2002Model.merge_profiles True >>> CosmoSac2002Model.temperature_exponents (1,)
Attributes
- num_segment_types#
Number of segment types.
Methods
- create_component(cosmo_string)[source]#
Create a
Componentconsistent with this model.- Parameters:
cosmo_string (str) – Contents of a COSMO output file.
- Returns:
Molecular component configured with the model’s parameters.
- Return type:
Examples
>>> from importlib.resources import files >>> from cosmolayer.cosmosac.model import CosmoSac2010Model >>> path = files("cosmolayer.data") / "C=C(N)O.cosmo" >>> component = CosmoSac2010Model.create_component(path.read_text()) >>> component.area 97.34554... >>> component.probabilities.shape (153,)
- create_interaction_matrices(temperature)[source]#
Create the interaction energy matrices at a given temperature.
- Parameters:
temperature (float) – Temperature in Kelvin.
- Returns:
Dimensionless interaction energy matrices ΔW/(RT). The number of matrices matches the length of
temperature_exponents.- Return type:
tuple[NDArray[np.float64], …]
Examples
>>> from cosmolayer.cosmosac import CosmoSac2002Model, CosmoSac2010Model
COSMO-SAC 2002 produces a single interaction matrix:
>>> matrices = CosmoSac2002Model.create_interaction_matrices(298.15) >>> len(matrices) 1 >>> matrix = matrices[0] >>> matrix.shape (51, 51) >>> print(matrix.min() < 0) # H-bonding can be favorable (negative) True >>> print(matrix.max() > 0) # Misfit interactions are unfavorable True
Plotting the COSMO-SAC 2002 interaction matrix:
>>> from cosmolayer.cosmosac import CosmoSac2002Model >>> from matplotlib import pyplot as plt >>> matrices = CosmoSac2002Model.create_interaction_matrices(298.15) >>> fig, ax = plt.subplots(figsize=(8, 6)) >>> im = ax.imshow(matrices[0], cmap="Spectral") >>> _ = fig.colorbar(im, ax=ax, label="ΔW/(RT)") >>> fig.tight_layout()
(
Source code,png,hires.png,pdf)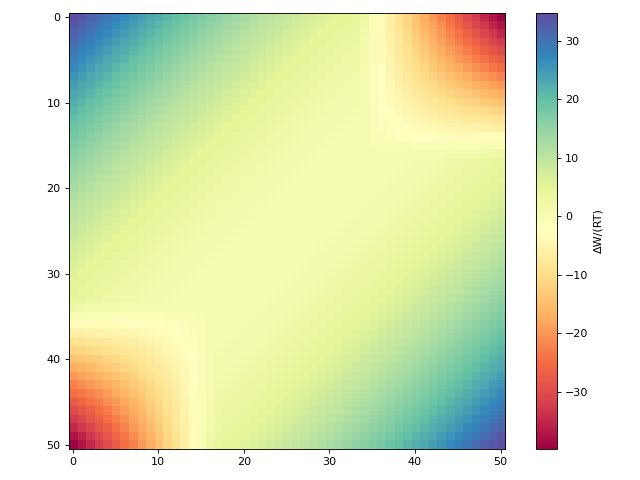
COSMO-SAC 2010 produces two matrices (one per temperature exponent):
>>> matrices = CosmoSac2010Model.create_interaction_matrices(298.15) >>> len(matrices) 2 >>> all(m.shape == (153, 153) for m in matrices) True
Plotting the COSMO-SAC 2010 interaction matrix:
>>> from cosmolayer.cosmosac import CosmoSac2010Model >>> from matplotlib import pyplot as plt >>> matrices = CosmoSac2010Model.create_interaction_matrices(298.15) >>> fig, ax = plt.subplots(figsize=(8, 6)) >>> im = ax.imshow(sum(matrices), cmap="Spectral") >>> _ = fig.colorbar(im, ax=ax, label="ΔW/(RT)") >>> fig.tight_layout()
(
Source code,png,hires.png,pdf)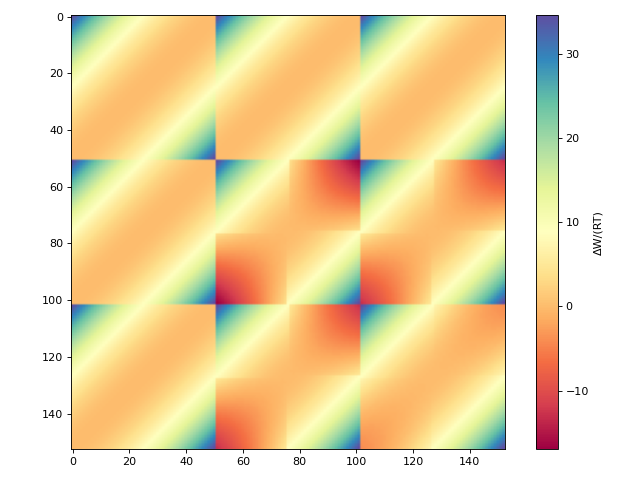
- create_mixture(components)[source]#
Create a
Mixtureconsistent with this model.- Parameters:
components (dict[str, str]) – Dictionary mapping component names to COSMO strings (contents of COSMO output files).
- Returns:
Mixture configured with the model’s parameters.
- Return type:
Examples
>>> from importlib.resources import files >>> from cosmolayer.cosmosac.model import CosmoSac2002Model, CosmoSac2010Model
Creating a mixture with the COSMO-SAC 2002 model:
>>> source = files("cosmolayer.data") >>> components = { ... "1-aminoethenol": (source / "C=C(N)O.cosmo").read_text(), ... "2-aminoethanol": (source / "NCCO.cosmo").read_text(), ... } >>> mixture = CosmoSac2002Model.create_mixture(components) >>> len(mixture) 2 >>> mixture.interaction_matrices(298.15)[0].shape (51, 51)
Creating a mixture with the COSMO-SAC 2010 model:
>>> mixture = CosmoSac2010Model.create_mixture(components) >>> len(mixture) 2 >>> matrices = mixture.interaction_matrices(298.15) >>> len(matrices) 2 >>> all(m.shape == (153, 153) for m in matrices) True